measure="PFT", the escalc() function automatically switches off the continuity correction.Note: Make sure you read the Note of Caution at the end of this page. In essence, I would strongly advise against using the Freeman-Tukey (double arcsine) transformation when meta-analyzing proportions.
When meta-analyzing proportions, it is usually advantageous to first transform the proportions into a measure that has better statistical properties (i.e., a sampling distribution that is closer to a normal distribution and whose sampling variance can be better approximated). An often recommended transformation in this context is the logit transformation (i.e., the log odds). While the sampling distribution of a logit transformed proportion is indeed better approximated by a normal distribution, the equation used to compute the corresponding sampling variance can still be quite inaccurate, especially when sample sizes are small. A transformation that works particularly well for normalizing and variance-stabilizing the sampling distribution of proportions is the Freeman-Tukey (double arcsine) transformation (Freeman & Tukey, 1950). The corresponding back-transformation equation was derived by Miller (1978).
The data used by Miller (1978) to illustrate the transformation and its inversion can be re-created with:
dat <- data.frame(xi=c(3, 6, 10, 1), ni=c(11, 17, 21, 6)) dat$pi <- with(dat, xi/ni) dat <- escalc(measure="PFT", xi=xi, ni=ni, data=dat)
The contents of dat are then:
xi ni pi yi vi 1 3 11 0.2727273 0.5695 0.0217 2 6 17 0.3529412 0.6444 0.0143 3 10 21 0.4761905 0.7626 0.0116 4 1 6 0.1666667 0.4758 0.0385
The yi values are the Freeman-Tukey (double arcsine) transformed proportions, while the vi values are the corresponding sampling variances.
Note: One can find two different definitions of the Freeman-Tukey transformation in the literature that differ only by the multiplicative constant $1/2$. The escalc() function includes the multiplicative constant $1/2$, while Miller (1978) leaves this out. Therefore, the transformed values given in the table by Miller are twice as large as the ones given above. Whether one includes the multiplicative constant or not is irrelevant, as long as one uses the correct equation for the sampling variance. For more details, see the question How is the Freeman-Tukey transformation of proportions and incidence rates computed? under the FAQ section.
We can check whether the back-transformation works for individual values with:
transf.ipft(dat$yi, dat$ni)
[1] 0.2727273 0.3529412 0.4761905 0.1666667
Those are indeed the individual proportions. Note that, due to the nature of the Freeman-Tukey transformation, the back-transformation requires information about the sample sizes of the individual studies. The relevance of this will become apparent in a moment.
As described by Miller (1978), we can aggregate the transformed values, either by computing an unweighted or a weighted mean (with inverse-variance weights). The unweighted mean can be obtained with:
res <- rma(yi, vi, method="EE", data=dat, weighted=FALSE) res
Equal-Effects Model (k = 4) I^2 (total heterogeneity / total variability): 0.00% H^2 (total variability / sampling variability): 0.73 Test for Heterogeneity: Q(df = 3) = 2.1754, p-val = 0.5368 Model Results: estimate se zval pval ci.lb ci.ub 0.6131 0.0734 8.3569 <.0001 0.4693 0.7569 *** --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
The value reported by Miller for the unweighted mean is again twice as large as the value given above, which we can confirm with:
predict(res, transf=function(x) x*2)
pred ci.lb ci.ub 1.2262 0.9386 1.5138
To back-transform the average, a value for the sample size is needed. Miller suggests to use the harmonic mean of the individual sample sizes in the inversion formula. This can be accomplished with:
predict(res, transf=transf.ipft.hm, targs=list(ni=dat$ni))
pred ci.lb ci.ub 0.3164 0.1794 0.4690
Therefore, the estimated true proportion based on the 4 studies is $.32$ (with 95% CI: $.18$ to $.47$).
Since the true proportions appear to be homogeneous (e.g., $Q(3) = 2.18$, $p=.54$), a more efficient estimate of the true proportion can be obtained by using inverse-variance weights. For this, we first synthesize the transformed values with:
res <- rma(yi, vi, method="EE", data=dat) res
Equal-Effects Model (k = 4) I^2 (total heterogeneity / total variability): 0.00% H^2 (total variability / sampling variability): 0.73 Test for Heterogeneity: Q(df = 3) = 2.1754, p-val = 0.5368 Model Results: estimate se zval pval ci.lb ci.ub 0.6547 0.0662 9.8853 <.0001 0.5249 0.7845 *** --- Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Again, the value reported by Miller is twice as large.
We can back-transform the estimated transformed average with:
predict(res, transf=transf.ipft.hm, targs=list(ni=dat$ni))
pred ci.lb ci.ub 0.3595 0.2297 0.4990
Therefore, the estimated true proportion is now equal to $.36$ (with 95% CI: $.23$ to $.50$).
When the event of interest is very rare or very common, the dataset may include proportions that are equal to 0 or 1. Such data can yield difficulties when meta-analyzing raw proportions (i.e., the sampling variance of such proportions will then be equal to 0) or when meta-analyzing logit transformed proportions (i.e., the log odds will then be either equal to minus or plus infinity). In practice, such cases are handled by adding a small constant (typically ½) to the data (sometimes called a 'continuity correction'). This approach may be acceptable when there are only a few such cases, but when meta-analyzing rare or common outcomes (where proportions equal to 0 or 1 are common), it is better to switch to a transformation that handles such proportions more gracefully, such as the Freeman-Tukey transformation. It does not require making any adjustments to the observed data, even when there are proportions equal to 0 or 1. As an example:
dat <- data.frame(xi=c(0,10), ni=c(10,10)) dat$pi <- with(dat, xi/ni) dat <- escalc(measure="PFT", xi=xi, ni=ni, data=dat, add=0)
yields the data frame:
xi ni pi yi vi 1 0 10 0 0.1531 0.0238 2 10 10 1 1.4177 0.0238
For both studies, we obtain a finite transformed value with a non-zero sampling variance. Note the use of add=0 to explicitly 'switch off' the default behavior of adding 1/2 to the data.1)
The back-transformation still works as intended:
transf.ipft(dat$yi, dat$ni)
[1] 0 1
When using the Freeman-Tukey transformation, an additional complication arises when using the back-transformation. The problem can be illustrated with some data:
dat <- data.frame(xi = c( 0, 4, 9, 16, 20), ni = c(10, 10, 15, 20, 20)) dat$pi <- with(dat, xi/ni) dat <- escalc(measure="PFT", xi=xi, ni=ni, data=dat, add=0)
xi ni pi yi vi 1 0 10 0.0 0.1531 0.0238 2 4 10 0.4 0.6936 0.0238 3 9 15 0.6 0.8799 0.0161 4 16 20 0.8 1.0901 0.0122 5 20 20 1.0 1.4608 0.0122
To back-transform the individual proportions, we need the individual sample sizes:
transf.ipft(dat$yi, dat$ni)
[1] 0.0 0.4 0.6 0.8 1.0
To back-transform the estimated average transformed proportion, we can use the harmonic mean of the sample sizes:
res <- rma(yi, vi, method="EE", data=dat) pred <- predict(res, transf=transf.ipft.hm, targs=list(ni=dat$ni)) pred
pred ci.lb ci.ub 0.6886 0.5734 0.7943
If we now want to draw a forest plot that includes not only the individual back-transformed proportions, but also the back-transformed average, things get complicated. If we use:
forest(res, transf=transf.ipft.hm, targs=list(ni=dat$ni), alim=c(0,1), refline=NA, digits=3)
then the resulting forest plot also uses the harmonic mean of the sample sizes for the back-transformation of the individual transformed proportions, which is not correct (resulting forest plot not shown).
We therefore need to first obtain the CI bounds of the individual studies with:
dat.back <- summary(dat, transf=transf.ipft, ni=dat$ni)
Now dat.back includes:
xi ni pi yi ci.lb ci.ub 1 0 10 0.0 0.0000 0.0000 0.1652 2 4 10 0.4 0.4000 0.1131 0.7238 3 9 15 0.6 0.6000 0.3382 0.8371 4 16 20 0.8 0.8000 0.5921 0.9513 5 20 20 1.0 1.0000 0.9157 1.0000
Here, the back-transformation is applied to each transformed proportion with the study-specific sample sizes. The yi values are therefore the back-transformed values (i.e., the raw proportions) and the ci.lb and ci.ub values are the back-transformed 95% CI bounds.
Finally, we can create the forest plot by directly passing the observed outcomes (i.e., proportions) and the CI bounds to the function. Then the back-transformed average with the corresponding CI bounds obtained earlier can be added to the plot with the addpoly() function. We add a couple tweaks to make the final forest plot look nice:
forest(dat.back$yi, ci.lb=dat.back$ci.lb, ci.ub=dat.back$ci.ub, psize=1, xlim=c(-0.5,1.8), alim=c(0,1), ylim=c(-2,8), refline=NA, digits=3L, xlab="Proportion", header=c("Study", "Proportion [95% CI]")) abline(h=0) addpoly(pred, row=-1, mlab="Equal-Effects Model")
The resulting figure then looks like this:
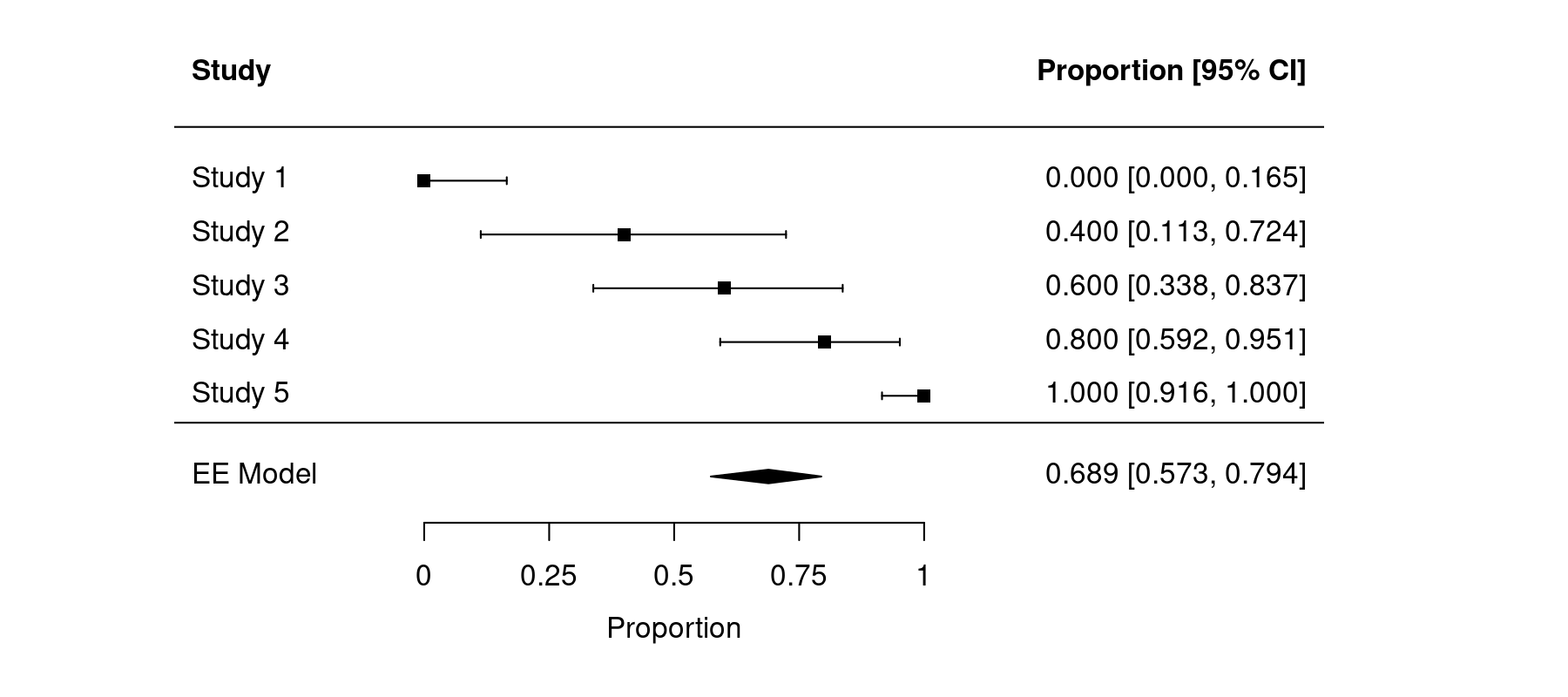
Now the estimated average and the results from the individual studies are correctly back-transformed.
It turns out that there are some major problems when using Freeman-Tukey (double arcsine) transformed proportions for a meta-analysis. For details, see Schwarzer et al. (2019) and Röver and Friede (2022). Therefore, I would strongly advise against using this transformation when meta-analyzing proportions. Alternatively, one can use the standard arcsine transformation (measure="PAS" in escalc()) or switch to a (random-effects) logistic regression model (which can be done with the rma.glmm() function).
Freeman, M. F., & Tukey, J. W. (1950). Transformations related to the angular and the square root. Annals of Mathematical Statistics, 21(4), 607–611. https://doi.org/10.1214/aoms/1177729756
Miller, J. J. (1978). The inverse of the Freeman-Tukey double arcsine transformation. American Statistician, 32(4), 138. https://doi.org/10.1080/00031305.1978.10479283
Röver, C., & Friede, T. (2022). Double arcsine transform not appropriate for meta‐analysis. Research Synthesis Methods, 13(5), 645–648. https://doi.org/10.1002/jrsm.1591
Schwarzer, G., Chemaitelly, H., Abu-Raddad, L. J. & Rücker, G. (2019). Seriously misleading results using inverse of Freeman-Tukey double arcsine transformation in meta-analysis of single proportions. Research Synthesis Methods, 10(3), 476–483. https://doi.org/10.1002/jrsm.1348
measure="PFT", the escalc() function automatically switches off the continuity correction.